A new study by 279 scientists, led by the Royal Botanic Gardens, Kew, has updated our understanding of the flowering plant tree of life by analyzing genetic data from over 9,500 species. This research, involving an international collaboration and significant technological advancements, provides crucial insights for plant classification, conservation, and medicinal discovery. The findings are freely accessible, promising to enhance future botanical studies and applications.
Scientists have constructed a groundbreaking tree of life using 1.8 billion letters of genetic code.
A recent study published in the journal Nature by an international team of 279 scientists, including three biologists from the University of Michigan, provides the latest insights into the flowering plant tree of life.
Using 1.8 billion letters of genetic code from more than 9,500 Kew’s Tree of Life Initiative.
“Analyzing this unprecedented amount of data to decode the information hidden in millions of DNA sequences was a huge challenge. But it also offered the unique opportunity to reevaluate and extend our knowledge of the plant tree of life, opening a new window to explore the complexity of plant evolution,” said Alexandre Zuntini, a research fellow at Royal Botanic Gardens, Kew.
Tom Carruthers, postdoctoral researcher in the lab of U-M evolutionary biologist Stephen Smith, is co-lead author of the study with Zuntini, who he previously worked with at Kew. U-M plant systematist Richard Rabeler is a co-author.
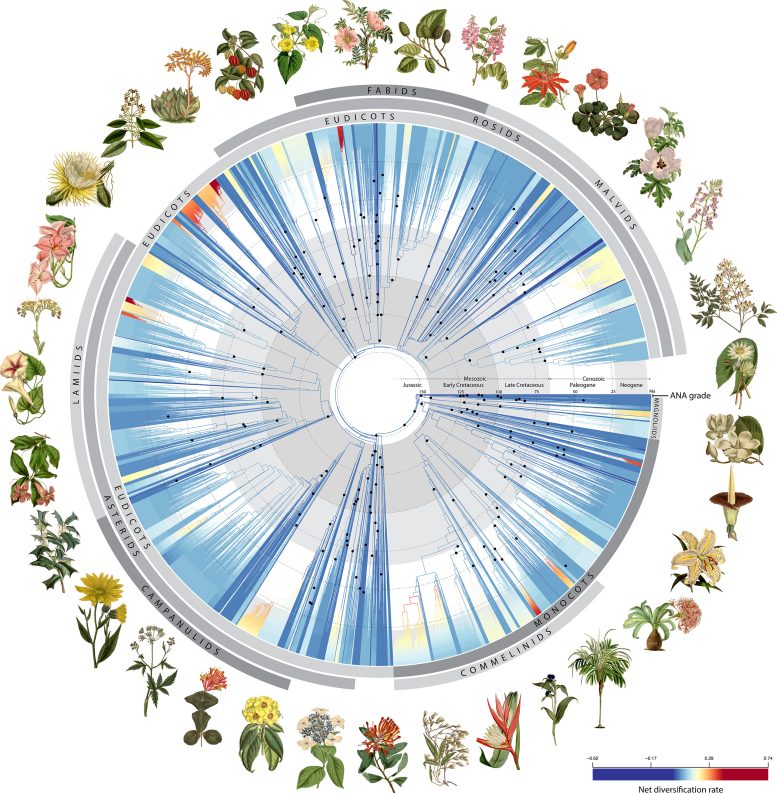
Angiosperm Tree of Life. Credit: RBG Kew
“Flowering plants feed, clothe, and greet us whenever we walk into the woods. The construction of a flowering plant tree of life has been a significant challenge and goal for the field of evolutionary biology for more than a century,” said Smith, co-author of the study and professor in the U-M Department of Ecology and Evolutionary Biology. “This project moves us closer to that goal by providing a massive dataset for most of the genera of flowering plants and offering one strategy to complete this goal.”
Smith had two roles on the project. First, members of his lab—including former U-M graduate student Drew Larson—traveled to Kew to help sequence members of a large and diverse plant group called Ericales, which includes blueberries, tea, ebony, azaleas, rhododendrons and Brazil nuts.
Second, Smith supervised the analyses and construction of the project dataset along with William Baker and Felix Forest of the Royal Botanic Gardens, Kew, and Wolf Eisenhardt of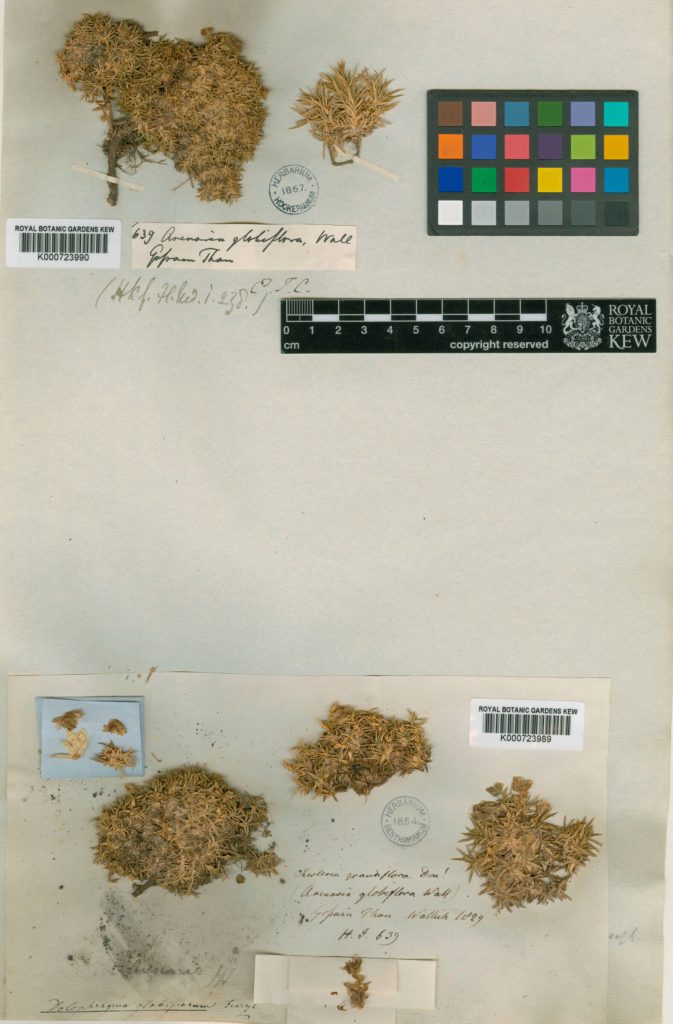 
Arenaria globilfora. Credit: RBG Kew
A key advantage of the team’s approach is that it enables a wide diversity of plant material, old and new, to be sequenced, even when the DNA is badly damaged. The vast treasure troves of dried plant material in the world’s herbarium collections, which comprise nearly 400 million scientific specimens of plants, can now be studied genetically.
“In many ways, this novel approach has allowed us to collaborate with the botanists of the past by tapping into the wealth of data locked up in historic herbarium specimens, some of which were collected as far back as the early 19th century,” said Baker, senior research leader for Kew’s Tree of Life Initiative.
“Our illustrious predecessors, such as Charles Darwin or Joseph Hooker, could not have anticipated how important these specimens would be in genomic research today. DNA was not even discovered in their lifetimes. Our work shows just how important these incredible botanical museums are to groundbreaking studies of life on Earth. Who knows what other undiscovered science opportunities lie within them?”
Across all 9,506 species sequenced, more than 3,400 came from material sourced from 163 herbaria in 48 countries.
“Sampling herbarium specimens for the study of plant relationships makes broad sampling from diverse areas of the world much more feasible than if one had to travel to get fresh material from the field,” said U-M’s Rabeler, a research scientist emeritus and former collection manager at the U-M Herbarium.
For the tree of life project, Rabeler helped verify the identity of herbarium specimens selected for sampling and analyzed the resulting data.
Flowering plants alone account for about 90% of all known plant life on land and are found virtually everywhere on the planet—from the steamiest tropics to the rocky outcrops of the Antarctic Peninsula. And yet, our understanding of how these plants came to dominate the scene soon after their origin has baffled scientists for generations, including Darwin.
Flowering plants originated more than 140 million years ago after which they rapidly overtook other vascular plants including their closest living relatives—the gymnosperms (nonflowering plants that have naked seeds, such as cycads, conifers, and ginkgo).
Darwin was mystified by the seemingly sudden appearance of such diversity in the fossil record. In an 1879 letter to Hooker, his close confidant and director of the Royal Botanic Gardens, Kew, he wrote: “The rapid development as far as we can judge of all the higher plants within recent geological times is an abominable mystery.”
Using 200 fossils, the authors scaled their tree of life to time, revealing how flowering plants evolved across geological time. They found that early flowering plants did indeed explode in diversity, giving rise to more than 80% of the major lineages that exist today shortly after their origin.
However, this trend then declined to a steadier rate for the next 100 million years until another surge in diversification about 40 million years ago, coinciding with a global decline in temperatures. These new insights would have fascinated Darwin and will surely help today’s scientists grappling with the challenges of understanding how and why species diversify.
Global Collaboration and Open Access
Assembling a tree of life this extensive would have been impossible without Kew’s scientists collaborating with many partners across the globe. In total, 279 authors were involved in the research, representing many different nationalities from 138 organizations in 27 countries.
“The plant community has a long history of collaborating and coordinating molecular sequencing to generate a more comprehensive and robust plant tree of life. The effort that led to this paper continues in that tradition but scales up quite significantly,” said U-M’s Smith.
The flowering plant tree of life has enormous potential in biodiversity research. This is because, just as one can predict the properties of an element based on its position in the periodic table, the location of a species in the tree of life allows us to predict its properties. The new data will thus be invaluable for enhancing many areas of science and beyond.
To enable this, the tree and all of the data that underpin it have been made openly and freely accessible to both the public and scientific community, including through theKew Tree of Life Explorer.
Open access will help scientists to make the best use of the data, such as combining it with artificial intelligence to predict which plant species may include molecules with medicinal potential.
Similarly, the tree of life can be used to better understand and predict how pests and diseases are going to affect plants in the future. Ultimately, the authors note, the applications of this data will be driven by the ingenuity of the scientists accessing it.
Reference: “Phylogenomics and the rise of the angiosperms” by Alexandre R. Zuntini, Tom Carruthers, Olivier Maurin, Paul C. Bailey, Kevin Leempoel, Grace E. Brewer, Niroshini Epitawalage, Elaine Françoso, Berta Gallego-Paramo, Catherine McGinnie, Raquel Negrão, Shyamali R. Roy, Lalita Simpson, Eduardo Toledo Romero, Vanessa M. A. Barber, Laura Botigué, James J. Clarkson, Robyn S. Cowan, Steven Dodsworth, Matthew G. Johnson, Jan T. Kim, Lisa Pokorny, Norman J. Wickett, Guilherme M. Antar, Lucinda DeBolt, Karime Gutierrez, Kasper P. Hendriks, Alina Hoewener, Ai-Qun Hu, Elizabeth M. Joyce, Izai A. B. S. Kikuchi, Isabel Larridon, Drew A. Larson, Elton John de Lírio, Jing-Xia Liu, Panagiota Malakasi, Natalia A. S. Przelomska, Toral Shah, Juan Viruel, Theodore R. Allnutt, Gabriel K. Ameka, Rose L. Andrew, Marc S. Appelhans, Montserrat Arista, María Jesús Ariza, Juan Arroyo, Watchara Arthan, Julien B. Bachelier, C. Donovan Bailey, Helen F. Barnes, Matthew D. Barrett, Russell L. Barrett, Randall J. Bayer, Michael J. Bayly, Ed Biffin, Nicky Biggs, Joanne L. Birch, Diego Bogarín, Renata Borosova, Alexander M. C. Bowles, Peter C. Boyce, Gemma L. C. Bramley, Marie Briggs, Linda Broadhurst, Gillian K. Brown, Jeremy J. Bruhl, Anne Bruneau, Sven Buerki, Edie Burns, Margaret Byrne, Stuart Cable, Ainsley Calladine, Martin W. Callmander, Ángela Cano, David J. Cantrill, Warren M. Cardinal-McTeague, Mónica M. Carlsen, Abigail J. A. Carruthers, Alejandra de Castro Mateo, Mark W. Chase, Lars W. Chatrou, Martin Cheek, Shilin Chen, Maarten J. M. Christenhusz, Pascal-Antoine Christin, Mark A. Clements, Skye C. Coffey, John G. Conran, Xavier Cornejo, Thomas L. P. Couvreur, Ian D. Cowie, Laszlo Csiba, Iain Darbyshire, Gerrit Davidse, Nina M. J. Davies, Aaron P. Davis, Kor-jent van Dijk, Stephen R. Downie, Marco F. Duretto, Melvin R. Duvall, Sara L. Edwards, Urs Eggli, Roy H. J. Erkens, Marcial Escudero, Manuel de la Estrella, Federico Fabriani, Michael F. Fay, Paola de L. Ferreira, Sarah Z. Ficinski, Rachael M. Fowler, Sue Frisby, Lin Fu, Tim Fulcher, Mercè Galbany-Casals, Elliot M. Gardner, Dmitry A. German, Augusto Giaretta, Marc Gibernau, Lynn J. Gillespie, Cynthia C. González, David J. Goyder, Sean W. Graham, Aurélie Grall, Laura Green, Bee F. Gunn, Diego G. Gutiérrez, Jan Hackel, Thomas Haevermans, Anna Haigh, Jocelyn C. Hall, Tony Hall, Melissa J. Harrison, Sebastian A. Hatt, Oriane Hidalgo, Trevor R. Hodkinson, Gareth D. Holmes, Helen C. F. Hopkins, Christopher J. Jackson, Shelley A. James, Richard W. Jobson, Gudrun Kadereit, Imalka M. Kahandawala, Kent Kainulainen, Masahiro Kato, Elizabeth A. Kellogg, Graham J. King, Beata Klejevskaja, Bente B. Klitgaard, Ronell R. Klopper, Sandra Knapp, Marcus A. Koch, James H. Leebens-Mack, Frederic Lens, Christine J. Leon, Étienne Léveillé-Bourret, Gwilym P. Lewis, De-Zhu Li, Lan Li, Sigrid Liede-Schumann, Tatyana Livshultz, David Lorence, Meng Lu, Patricia Lu-Irving, Jaquelini Luber, Eve J. Lucas, Manuel Luján, Mabel Lum, Terry D. Macfarlane, Carlos Magdalena, Vidal F. Mansano, Lizo E. Masters, Simon J. Mayo, Kristina McColl, Angela J. McDonnell, Andrew E. McDougall, Todd G. B. McLay, Hannah McPherson, Rosa I. Meneses, Vincent S. F. T. Merckx, Fabián A. Michelangeli, John D. Mitchell, Alexandre K. Monro, Michael J. Moore, Taryn L. Mueller, Klaus Mummenhoff, Jérôme Munzinger, Priscilla Muriel, Daniel J. Murphy, Katharina Nargar, Lars Nauheimer, Francis J. Nge, Reto Nyffeler, Andrés Orejuela, Edgardo M. Ortiz, Luis Palazzesi, Ariane Luna Peixoto, Susan K. Pell, Jaume Pellicer, Darin S. Penneys, Oscar A. Perez-Escobar, Claes Persson, Marc Pignal, Yohan Pillon, José R. Pirani, Gregory M. Plunkett, Robyn F. Powell, Ghillean T. Prance, Carmen Puglisi, Ming Qin, Richard K. Rabeler, Paul E. J. Rees, Matthew Renner, Eric H. Roalson, Michele Rodda, Zachary S. Rogers, Saba Rokni, Rolf Rutishauser, Miguel F. de Salas, Hanno Schaefer, Rowan J. Schley, Alexander Schmidt-Lebuhn, Alison Shapcott, Ihsan Al-Shehbaz, Kelly A. Shepherd, Mark P. Simmons, André O. Simões, Ana Rita G. Simões, Michelle Siros, Eric C. Smidt, James F. Smith, Neil Snow, Douglas E. Soltis, Pamela S. Soltis, Robert J. Soreng, Cynthia A. Sothers, Julian R. Starr, Peter F. Stevens, Shannon C. K. Straub, Lena Struwe, Jennifer M. Taylor, Ian R. H. Telford, Andrew H. Thornhill, Ifeanna Tooth, Anna Trias-Blasi, Frank Udovicic, Timothy M. A. Utteridge, Jose C. Del Valle, G. Anthony Verboom, Helen P. Vonow, Maria S. Vorontsova, Jurriaan M. de Vos, Noor Al-Wattar, Michelle Waycott, Cassiano A. D. Welker, Adam J. White, Jan J. Wieringa, Luis T. Williamson, Trevor C. Wilson, Sin Yeng Wong, Lisa A. Woods, Roseina Woods, Stuart Worboys, Martin Xanthos, Ya Yang, Yu-Xiao Zhang, Meng-Yuan Zhou, Sue Zmarzty, Fernando O. Zuloaga, Alexandre Antonelli, Sidonie Bellot, Darren M. Crayn, Olwen M. Grace, Paul J. Kersey, Ilia J. Leitch, Hervé Sauquet, Stephen A. Smith, Wolf L. Eiserhardt, Félix Forest and William J. Baker, 24 April 2024, Nature.
DOI: 10.1038/s41586-024-07324-0