By

A groundbreaking study employing full genome-scale data has revolutionized our understanding of bird evolution, revealing new insights into the phylogenetic relationships among bird species. By analyzing a comprehensive dataset of 363 bird species, the research has redefined the avian family tree, identifying four major clades within Neoaves and shedding light on the complex evolutionary history of birds. Credit: SciTechDaily.com
The most extensive genomic study to date has unveiled how the birds spread all over the world after mass extinction.
Birds represent the sole surviving lineage of dinosaurs in the present day. Roughly 66 million years ago, at the transition from the 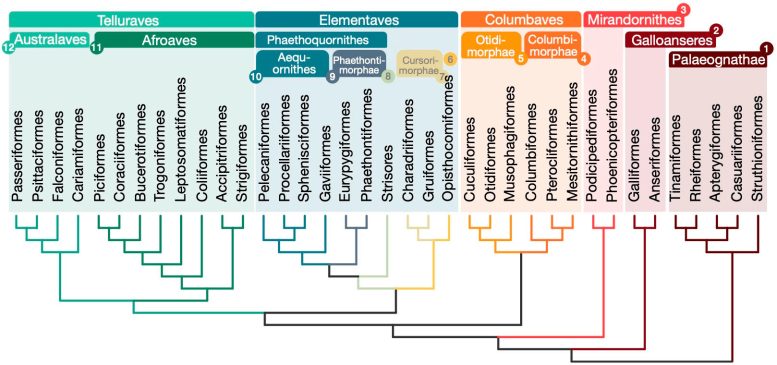
Relationships for 363 bird species based on 63,430 intergenic loci. Credit: Josefin Stiller
The new tree is challenging the organization of Neoaves by classifying this big group into four major clades: Mirandornithes, Columbaves, Elementaves, and Telluraves. Elementaves is a newly proposed grouping comprising ca. 14% of all species of modern birds including disparate groups such as the enigmatic hoatzin, shorebirds, hummingbirds, and tropicbirds.
Elementaves is named to reflect the group’s remarkable diversity in ecological niches, representing the major elements of Earth, air, and water. This new family tree resolves some long-standing debates over the relationships among avian species and lays a solid foundation for studying avian evolution and trait development.
The Impact of Comprehensive Genomic Analysis
By employing full genome data across 363 bird species, this is the largest-ever dataset used for phylogenetic analyses of birds. The team built a new pipeline to extract over 150,000 regions spread out across the genome.
“We characterized phylogenetic relationships across the entire genome and identified patterns associated with the genomic context and sequence characteristics,” said Josefin Stiller, the leading author of this study and an Assistant Professor in evolutionary biology at the University of Copenhagen, “We found that various parts of the genome, for example, individual chromosomes or protein-coding genes, often support vastly different trees. This likely explains why studies that only analyzed certain genomic parts were in conflict.”
Putting together high-quality and large amounts of data was key to producing a robust phylogenetic tree. The team discovered that for most branches, a consensus on their relationships can be reached when a sufficient amount of data was used. But the phylogenetic positions for some bird groups like owls and hawks remain puzzling even with a full-genome scale of data.
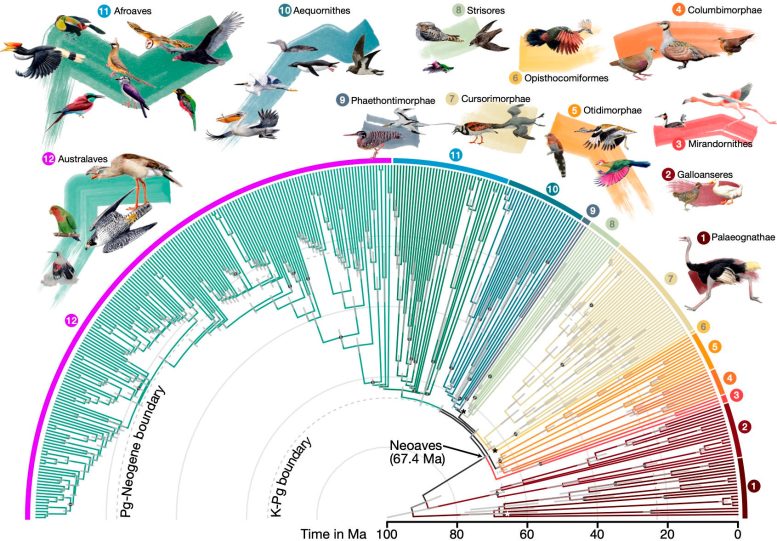
Divergence times for 363 bird species based on 63,430 intergenic loci. Credit: Josefin Stiller
“More data does not necessarily produce a better solution,” Zhang said. Siavash Mirarab, co-senior author of the study, a professor of electrical and computer engineering at the University of California, San Diego, added that, “The reason for this may be some complex evolutionary history like ancient cross-mating between two lineages, incomplete lineage sorting, long-branch attraction, and biased DOI: 10.1038/s41586-024-07323-1


