
The mapping of the sugarcane genome marks a scientific milestone, unlocking new possibilities in crop improvement and disease management, thus fostering advances in agricultural productivity and bioenergy development. Credit: SciTechDaily.com
Scientists created a highly accurate reference genome for one of the most important modern crops and found a rare example of how genes confer disease resistance in plants. Exploring sugarcane’s genetic code could help researchers develop more resilient and productive crops, with implications for both sugar production and biofuels.
- Until now, sugarcane’s complicated genetics made it the last major crop without a complete and highly accurate genome.
- Researchers combined multiple techniques to successfully map out sugarcane’s Phytozome.
“This was the most complicated genome sequence we’ve yet completed,” said Jeremy Schmutz, Plant Program lead at the JGI and faculty investigator at the HudsonAlpha Institute for Biotechnology. “It shows how far we’ve come. This is the kind of thing that 10 years ago people thought was impossible. We’re able to accomplish goals now that we just didn’t think were possible to do in plant genomics.”
Sugarcane’s genome is so complex both because it is large and because it contains more copies of chromosomes than a typical plant, a feature called polyploidy. Sugarcane has about 10 billion base pairs, the building blocks of DNA; for comparison, the human genome has about 3 billion.
Many sections of sugarcane’s DNA are identical both within and across different chromosomes. That makes it a challenge to correctly reassemble all the small segments of DNA while reconstructing the full genetic blueprint. Researchers solved the puzzle by combining multiple genetic sequencing techniques, including a newly developed method known as PacBio HiFi sequencing that can accurately determine the sequence of longer sections of DNA.
Understanding and Utilizing the Sugarcane Genome
Having a complete “reference genome” makes it easier to study sugarcane, enabling researchers to compare its genes and pathways with those in other well-studied crops such as sorghum or other biofuel crops of interest, like switchgrass and miscanthus. By comparing this reference to other crops, it becomes easier to understand how each gene influences a trait of interest, such as which genes are highly expressed during sugar production, or which genes are important for disease resistance. This study found that the genes responsible for resistance to brown rust, a fungal pathogen that previously caused millions of dollars of damage to sugarcane crops, are found in only one location in the genome.
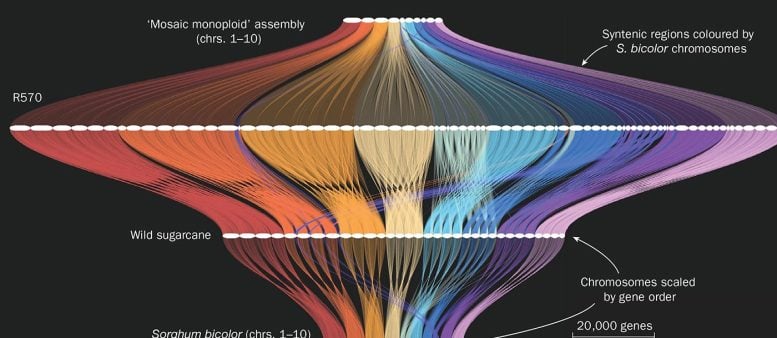
This image shows a gene order map (created using GENESPACE) that compares genome assemblies among related plant species. The horizontal white lines represent chromosomes, and the colored braids that link them show conserved blocks of genes. This enables researchers to track conserved genes of interest from well-researched crops (such as Sorghum bicolor; a specific type of sorghum) into more complex genomes, such as wild sugarcane and cultivar R570, to better understand their function. For contrast, the previous monoploid assembly of R570 is provided on the top row, where multiple chromosome copies in the genome were represented as a single, mosaic assembly. Credit: Adam Healey and John Lovell/HudsonAlpha
“When we sequenced the genome, we were able to fill a gap in the genetic sequence around brown rust disease,” said Adam Healey, first author of the paper and a researcher at HudsonAlpha.
“There are hundreds of thousands of genes in the sugarcane genome, but it’s only two genes, working together, that protect the plant from this pathogen. Across plants, there are only a handful of instances that we know of where protection works in a similar way. Better understanding of how this disease resistance works in sugarcane could help protect other crops facing similar pathogens down the road.”
Researchers studied a cultivar of sugarcane known as R570 that has been used for decades around the world as the model to understand sugarcane genetics. Like all modern sugarcane cultivars, R570 is a hybrid made by crossing the domesticated DOI: 10.1038/s41586-024-07231-4
This study involved collaborations with institutes from around the world, including France (CIRAD, UMR-AGAP, ERCANE); Australia (


